Anti-Phospho-ATM-Ser-1981 antibody [M366] (STJA0041054)
SPECIFICATIONS
ClonalityMonoclonal
HostMouse
ConjugationUnconjugated
IsotypeIgG1
ImmunogenClone M366 was generated from a phospho-peptide that included amino acids surrounding Serine 1981 in human ATM. This sequence has high homology to the conserved site in rat and mouse ATM.
General Information
| Short Description | Mouse monoclonal anti-Phospho-ATM-Ser-1981 for use in ELISA, WB, IP and ICC in Human, Mouse and Rat samples. Datasheet included with dilution recommendations, and related reagents. |
| Applications | ELISA/WB/IP/ICC |
| Host | Mouse |
| Reactivity | Human/Mouse/Rat |
| Note | STRICTLY FOR FURTHER SCIENTIFIC RESEARCH USE ONLY (RUO). MUST NOT TO BE USED IN DIAGNOSTIC OR THERAPEUTIC APPLICATIONS. |
Product Properties
| Clonality | Monoclonal |
| Clone ID | M366 |
| Isotype | IgG1 |
| Conjugation | Unconjugated |
| Purification | Protein A Purified |
| Dilution Range | WB 1:1000ICC 1:200IP 1:100 |
| Formulation | PBS + 1 mg/ml BSA, 0.05% NaN3 and 50% glycerol |
| Storage Instruction | Storage at-20°C is recommended, as aliquots may be taken without freeze/thawing due to presence of 50% glycerol. Stable for at least 1 year at-20°C. |
Target Information
| Gene Symbol | ATM |
| Gene ID | 472 |
| Uniprot ID | ATM_HUMAN |
| Immunogen | Clone M366 was generated from a phospho-peptide that included amino acids surrounding Serine 1981 in human ATM. This sequence has high homology to the conserved site in rat and mouse ATM. |
| Immunogen Sequence | Human |
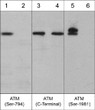
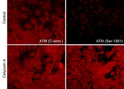
| Specificity | This antibody detects a 370 kDa* protein corresponding to the molecular mass of ATM on SDS-PAGE immunoblots of calyculin A treated human A431 and Jurkat Cells, but is not observed in control cells. |
Additional Info
| Tissue Specificity | Found in pancreas, kidney, skeletal muscle, liver, lung, placenta, brain, heart, spleen, thymus, testis, ovary, small intestine, colon and leukocytes. |
| Post Translational Modifications | Phosphorylated by NUAK1/ARK5. Autophosphorylation on Ser-367, Ser-1893, Ser-1981 correlates with DNA damage-mediated activation of the kinase. During the late stages of DNA damage response, dephosphorylated following deacetylation by SIRT7, leading to ATM deactivation. Acetylation, on DNA damage, is required for activation of the kinase activity, dimer-monomer transition, and subsequent autophosphorylation on Ser-1981. Acetylated in vitro by KAT5/TIP60. Deacetylated by SIRT7 during the late stages of DNA damage response, promoting ATM dephosphorylation and subsequent deactivation. |
| Function | Serine/threonine protein kinase which activates checkpoint signaling upon double strand breaks (DSBs), apoptosis and genotoxic stresses such as ionizing ultraviolet A light (UVA), thereby acting as a DNA damage sensor. Recognizes the substrate consensus sequence ST-Q. Phosphorylates 'Ser-139' of histone variant H2AX at double strand breaks (DSBs), thereby regulating DNA damage response mechanism. Also plays a role in pre-B cell allelic exclusion, a process leading to expression of a single immunoglobulin heavy chain allele to enforce clonality and monospecific recognition by the B-cell antigen receptor (BCR) expressed on individual B-lymphocytes. After the introduction of DNA breaks by the RAG complex on one immunoglobulin allele, acts by mediating a repositioning of the second allele to pericentromeric heterochromatin, preventing accessibility to the RAG complex and recombination of the second allele. Also involved in signal transduction and cell cycle control. May function as a tumor suppressor. Necessary for activation of ABL1 and SAPK. Phosphorylates DYRK2, CHEK2, p53/TP53, FBXW7, FANCD2, NFKBIA, BRCA1, CREBBP/CBP, RBBP8/CTIP, FBXO46, MRE11, nibrin (NBN), RAD50, RAD17, PELI1, TERF1, UFL1, RAD9, UBQLN4 and DCLRE1C. May play a role in vesicle and/or protein transport. Could play a role in T-cell development, gonad and neurological function. Plays a role in replication-dependent histone mRNA degradation. Binds DNA ends. Phosphorylation of DYRK2 in nucleus in response to genotoxic stress prevents its MDM2-mediated ubiquitination and subsequent proteasome degradation. Phosphorylates ATF2 which stimulates its function in DNA damage response. Phosphorylates ERCC6 which is essential for its chromatin remodeling activity at DNA double-strand breaks. Phosphorylates TTC5/STRAP at 'Ser-203' in the cytoplasm in response to DNA damage, which promotes TTC5/STRAP nuclear localization. Also involved in pexophagy by mediating phosphorylation of PEX5: translocated to peroxisomes in response to reactive oxygen species (ROS), and catalyzes phosphorylation of PEX5, promoting PEX5 ubiquitination and induction of pexophagy. |
| Protein Name | Serine-Protein Kinase AtmAtaxia Telangiectasia MutatedA-T Mutated |
| Database Links | Reactome: R-HSA-2559586Reactome: R-HSA-3371453Reactome: R-HSA-349425Reactome: R-HSA-5685938Reactome: R-HSA-5685942Reactome: R-HSA-5693548Reactome: R-HSA-5693554Reactome: R-HSA-5693565Reactome: R-HSA-5693568Reactome: R-HSA-5693571Reactome: R-HSA-5693579Reactome: R-HSA-5693607Reactome: R-HSA-5693616Reactome: R-HSA-6796648Reactome: R-HSA-6803204Reactome: R-HSA-6803207Reactome: R-HSA-6804756Reactome: R-HSA-6804757Reactome: R-HSA-6804760Reactome: R-HSA-69473Reactome: R-HSA-69541Reactome: R-HSA-912446Reactome: R-HSA-9664873Reactome: R-HSA-9701192Reactome: R-HSA-9704331Reactome: R-HSA-9704646Reactome: R-HSA-9709570Reactome: R-HSA-9709603 |
| Cellular Localisation | NucleusCytoplasmic VesicleCytoplasmCytoskeletonMicrotubule Organizing CenterCentrosomePeroxisome MatrixPrimarily NuclearFound Also In Endocytic Vesicles In Association With Beta-AdaptinTranslocated To Peroxisomes In Response To Reactive Oxygen Species (Ros) By Pex5 |
| Alternative Antibody Names | Anti-Serine-Protein Kinase Atm antibodyAnti-Ataxia Telangiectasia Mutated antibodyAnti-A-T Mutated antibodyAnti-ATM antibody |
Information sourced from Uniprot.org