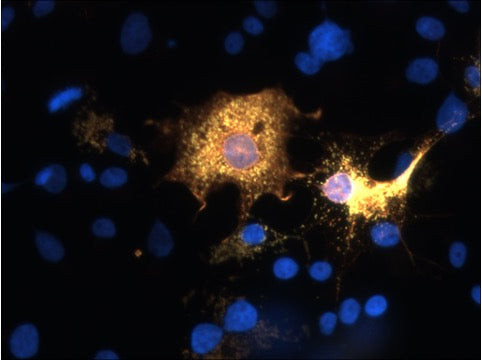
| Tissue Specificity | Widely expressed in the brain and neurons (at protein level). Expressed in photoreceptor cells of the neural retina. |
| Post Translational Modifications | Phosphorylation at Ser-548 by JNK kinases (MAPK8, MAPK9 and /or MAPK10) enhance the NAD(+) hydrolase (NADase) activity. Phosphorylation at Ser-548 and subsequent activation takes place in response to oxidative stress conditions and inhibits mitochondrial respiration. |
| Function | NAD(+) hydrolase, which plays a key role in axonal degeneration following injury by regulating NAD(+) metabolism. Acts as a negative regulator of MYD88- and TRIF-dependent toll-like receptor signaling pathway by promoting Wallerian degeneration, an injury-induced form of programmed subcellular death which involves degeneration of an axon distal to the injury site. Wallerian degeneration is triggered by NAD(+) depletion: in response to injury, SARM1 is activated and catalyzes cleavage of NAD(+) into ADP-D-ribose (ADPR), cyclic ADPR (cADPR) and nicotinamide.NAD(+) cleavage promoting cytoskeletal degradation and axon destruction. Also able to hydrolyze NADP(+), but not other NAD(+)-related molecules. Can activate neuronal cell death in response to stress. Regulates dendritic arborization through the MAPK4-JNK pathway. Involved in innate immune response: inhibits both TICAM1/TRIF- and MYD88-dependent activation of JUN/AP-1, TRIF-dependent activation of NF-kappa-B and IRF3, and the phosphorylation of MAPK14/p38. |
| Protein Name | Nad(+ Hydrolase Sarm1Nadase Sarm1Nadp(+ Hydrolase Sarm1Sterile Alpha And Tir Motif-Containing Protein 1 |
| Database Links | Reactome: R-MMU-166166Reactome: -MMU-936964Reactome: -MMU-937041Reactome: -MMU-937072 |
| Cellular Localisation | CytoplasmCell ProjectionAxonDendriteSynapseMitochondrionAssociated With Microtubules |
| Alternative Antibody Names | Anti-Nad(+ Hydrolase Sarm1 antibodyAnti-Nadase Sarm1 antibodyAnti-Nadp(+ Hydrolase Sarm1 antibodyAnti-Sterile Alpha And Tir Motif-Containing Protein 1 antibodyAnti-Sarm1 antibodyAnti-Kiaa0524 antibody |
Information sourced from Uniprot.org